DOE and the HGP
Why was the Department of Energy involved in the Human Genome Project?
After the atomic bomb was developed and used, the U.S. Congress charged the U.S. Department of Energy’s (DOE) predecessor agencies (the Atomic Energy Commission and the Energy Research and Development Administration) with studying and analyzing genome structure, replication, damage, and repair and the consequences of genetic mutations, especially those caused by radiation and chemical by-products of energy production. From these studies grew the recognition that the best way to study these effects was to analyze the entire human genome to obtain a reference sequence.
Planning began in 1986 for DOE’s Human Genome Program and in 1987 for the National Institutes of Health’s (NIH) program. The DOE-NIH U.S. Human Genome Project formally began October 1, 1990, after the first joint 5-year plan was written and a memorandum of understanding was signed between the two organizations. For more information see History of the Human Genome Project and DOE Biological and Environmental Research Program.
Consistent with the goals of the Human Genome Project, the DOE Human Genome Program focused on the following:
- Mapping (more information) human chromosomes 2, 5, 11, X, 16, 19, and 21;
- Comparative studies between mouse and human genomes;
- Development of important biological resources for the Human Genome Project and the broader biomedical research communities, including purified DNA collections for each human chromosome and sequence-ready DNA;
- Technologies, instrumentation, and robotics for more efficient DNA sequencing;
- Development of analysis algorithms and integration of databases (informatics) for managing and interpreting genome data; and
- Communicating about the Human Genome Project to those who would interpret it for various professions and ultimately for the public.
Another important DOE goal was to foster research into the ethical, legal, and social implications (ELSI) of genome research. The DOE Human Genome Program ELSI component and the data it generated concentrated on two main areas: (1) privacy and confidentiality of personal genetic information, including its accumulation in large, computerized databases and databanks; and (2) development of educational materials and activities in genome science and ELSI, including curricula and TV documentaries, workshops, and seminars for targeted audiences. Other areas of interest include data privacy arising from potential uses of genetic testing in the workplace and issues related to commercialization of genome research results and technology transfer.
For more details on the Department of Energy’s involvement, see the following:
- “Evolution of a Vision — Part I” by David Smith
- “Evolution of a Vision — Part II” by Francis S. Collins
- “The Alta Summit, December 1984” by Robert Mullan Cook-Deegan
What DOE investments improved the efficiency of the Human Genome Project by reducing costs, speeding progress, and furthering technology?
Making the Project Possible
Its long-standing mission to understand and characterize the potential health risks posed by energy use and production led DOE to propose, in the mid-1980s, that all three billion bases of DNA from an “average” human should be sequenced. Technologies available before that time had not enabled the routine detection of extremely rare and often minute genetic changes resulting from radiation and chemical exposures.
The scientific foundation for DOE’s Human Genome Initiative already existed at the national laboratories.
- DOE had a long history of conducting large multidisciplinary projects involving biologists, chemists, engineers, and mathematicians.
- Genbank, a DNA sequence repository, had been developed at Los Alamos National Laboratory (LANL) with DOE computer and data-management expertise. Today, Genbank, the world’s principal DNA sequence database, resides at the National Library of Medicine.
- Chromosome-sorting capabilities essential to a genome initiative existed at LANL and Lawrence Livermore National Laboratory (LLNL). Using this technology, LANL and LLNL began the National Laboratory Gene Library Project, a collection of cloned DNAs from single human chromosomes.
In 1986, DOE became the first federal agency to announce and fund a genome program.
Developing the Tools and Technologies for Success
[NOTE: The DOE investments described below helped make the Human Genome Project a success. Substantial investments by NIH and the Wellcome Trust in the United Kingdom were equally important, however, and should not be overlooked. In most cases, the DOE successes outlined below were the result of basic research programs. Research is an incremental process that learns from both the successes and failures of other research investments, including those at other agencies and organizations. In addition, no single instrument, technology, reagent, or protocol made high-throughput DNA sequencing possible, many contributors were responsible.]
- DNA Sequencers: Research on capillary-based DNA sequencing contributed to the development of the two major DNA sequencing machines—the Perkin-Elmer 3700 and the MegaBace DNA sequencers. The MegaBace DNA sequencer was developed initially with DOE funds by Dr. Richard Mathies at U.C. Berkeley. The Perkin-Elmer 3700 was based, in part, on DOE-funded research by Dr. Norman Dovichi at the University of Alberta. These high-throughput instruments are one of the keys to the success of the genome project.
- Fluorescent dyes: DNA sequencing originally used radiolabeled DNA subunits. DOE-funded research contributed to the development of fluorescent dyes that increased the accuracy and safety of DNA sequencing as well as the ability to automate the procedures.
- DNA cloning vectors: Before large DNA molecules can be sequenced, they are cut into small pieces and multiplied, or cloned, into numerous copies using microbial-based “cloning” vectors. Today, the bacterial artificial chromosome (BAC) is the most commonly used vector for initial DNA amplification before sequencing. These cloning vectors were developed with DOE funds.
- BAC-end sequencing: The widely agreed-upon strategy for sequencing the human genome is based on the use of BACs that carry fragments of human DNA from known locations in the genome. DOE-funded research at The Institute for Genomic Research in Rockville, Maryland, and at the University of Washington provided the sequencing community with a complete set of over 450,000 BAC-based genetic “markers” corresponding to a sequence tag every 3 to 4 kilobases across the entire human genome. These markers were needed to assemble both the draft and the final human DNA sequence.
- GRAIL: Gene Recognition and Assembly Internet Link (GRAIL) is one of the most widely used computer programs for identifying potential genes in DNA sequence and for general DNA sequence analysis. This powerful analytical tool was developed with DOE funds by Dr. Ed Uberbacher at Oak Ridge National Laboratory. Although a number of gene-finding tools are now available for use, GRAIL led the way.
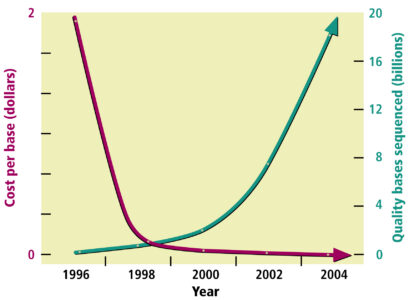
Reducing Costs and Speeding Up Sequencing
The above technological developments dramatically decreased DNA sequencing’s cost while increasing its speed and efficiency. For example, it took 4 years for the international Human Genome Project to produce the first billion base pairs of sequence and less than 4 months to produce the second billion base pairs. In the month of January 2003, the DOE team sequenced 1.5 billion bases. The cost of sequencing has dropped dramatically since the project began and is still dropping rapidly.
